Abstract
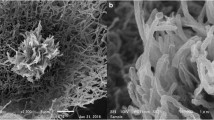
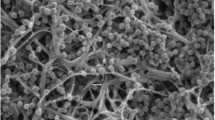
Myceliophthora guttulata sp. nov. is described and illustrated based on strains isolated from soil in China. This species is thermophilic with optimal growth temperature of 40–45 °C, and minimum growth temperature of 25 °C. Morphologically, this species is characterized by smooth, guttulate, pyriform to obovoid blastoconidia born directly on the side of hyphae, on long or short pedicels or in groups of 1–4 on ampulliform swellings. Phylogenetic analysis based on multi-locus alignment of internal transcribed spacer (ITS), elongation factor 1-alpha (EF1-α) and RNA polymerase II subunit (RPB2) regions showed that M. guttulata clustered within the genus Myceliophthora, and is closely related to four thermophilic species, i.e. M. fergusii, M. thermophila, M. heterothallic, and M. hinnulea.


Similar content being viewed by others
References
Awao T, Udagawa SI (1983) A new thermophilic species of Myceliophthora. Mycotaxon 16:436–440
Basu M (1985) Myceliophthora indica in a new thermophilic species from India. Nova Hedwig 40:85–90
Cai L, Jeewon R, Hyde KD (2006) Phylogenetic investigations of Sordariaceae based on multiple gene sequences and morphology. Mycol Res 110:137–150
Carmichael JW (1962) Chrysosporium and some other aleuriosporic hyphomycetes. Can J Bot 40:1137–1173
Cooney DG, Emerson R (1964) Thermophilic fungi-An account of their biology, activities and classification. Freeman, San Francisco
Costantin J (1892) Sur quelques maladies du blanc de champignons. Cr Hebd Séanc Acad Sci Paris 114:849–851
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Emmons CW (1932) The development of the ascocarp in two species of Thielavia. B Torrey Bot Club 59:415–422
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Huelsenbeck JP, Ronquist F (2001) Mrbayes: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Maheshwari R, Bharadwaj G, Bhat MK (2000) Thermophilic fungi: their physiology and enzymes. Microbiol Mol Biol R 64:461–488
Mouchacca J (1997) Thermophilic fungi: biodiversity and taxonomic status. Cryptogamie Mycol 18:19–69
Nylander JAA (2004) MrModeltest v2. Program distributed by the author. Evolutionary Biology Centre, Uppsala University
Page RDM (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput App Biosci 12:357–358
Rannala B, Yang Z (1996) Probability distribution of molecular evolutionary trees: a new method of phylogenetic inference. J Mol Evol 43:304–311
Shimodaira H, Hasegawa M (1999) Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol Biol Evol 16:1114–1116
Stchigel AM, Sagués M, Cano J, Guarro J (2000) Three new thermotolerant species of Corynascus from soil, with a key to the known species. Mycol Res 104:879–887
Swofford DL (2003) PAUP*: phylogenetic analysis using parsimony, version 4.0 b10. Sinauer, Sunderland
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Udagawa SH, Horie F (1972) A new species of Thielavia and its Chrysosporium conidial state. Bull Natl Sci Mus Tokyo 15:191–196
van den Brink J, Samson RA, Hagen F, Boekhout T, de Vries RP (2012) Phylogeny of the industrial relevant, thermophilic genera Myceliophthora and Corynascus. Fungal Divers 52:197–207
van Oorschot CAN (1977) The genus Myceliophthora. Persoonia 9:401–408
van Oorschot CAN (1980) A revision of Chrysosporium and allied genera. Stud Mycol 20:1–89
von Arx JA (1973) Further observations on Sporotrichum and some similar fungi. Persoonia 7:127–130
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: a guide to methods and applications. Academic Press, pp 315–322
Zhaxybayeva O, Gogarten JP (2002) Bootstrap, Bayesian probability and maximum likelihood mapping: exploring new tools for comparative genome analyses. Genomics 3:1–15
Acknowledgments
This study was financially supported by NSFC 31110103906 and CASKSCX2-YW-Z-1026. Wei Sun and Fang Liu are thanked for giving technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, Y., Wu, WP., Hu, DM. et al. A new thermophilic species of Myceliophthora from China. Mycol Progress 13, 165–170 (2014). https://doi.org/10.1007/s11557-013-0904-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-013-0904-8