Abstract
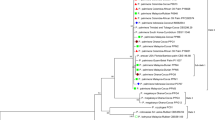
Sarocladium oryzae is a common plant pathogen causing rice sheath rot worldwide. Sarocladium attenuatum has been identified as one of the causal agents of rice sheath rot and was synonymized under S. oryzae. The analysis of 102 Sarocladium strains associated with rice in Taiwan, based on morphological features and multilocus molecular phylogeny, reveals that these two species are distinct and that two other species, S. sparsum sp. nov. and S. spirale sp. nov., can be identified. Sarocladium sparsum is phylogenetically closely related to S. attenuatum and S. oryzae and causes the same disease symptom in pathogenicity tests. Sarocladium spirale, a sister taxon of S. hominis and S. spinificis, is not pathogenic to rice. All strains of S. attenuatum and some S. sparsum strains show fluorescence emission ability under 365-nm UV light, while S. oryzae does not.







Similar content being viewed by others
References
Bills GF, Platas G, Gams W (2004) Conspecificity of the cerulenin and helvolic acid producing ‘Cephalosporium caerulens’, and the hypocrealean fungus Sarocladium oryzae. Mycol Res 108:1291–1300. https://doi.org/10.1017/S0953756204001297
Brady BLK (1980) Sarocladium attenuatum. CMI Descriptions of Pathogenic Fungi and Bacteria No. 673
Bridge PD, Hawksworth DL, Kavishe DF et al (1989) A revision of the species concept in Sarocladium, the causal agent of sheath-rot in rice and bamboo blight, based on biochemical and morphometric analyses. Plant Pathol 38:239–245. https://doi.org/10.1111/j.1365-3059.1989.tb02139.x
Chen JD, Zhang GC, Fu XH (1986) The pathogen of rice purple sheath disease Sarocladium sinense sp. nov. Acta Mycol Sin Suppl 1:318–327
Crous PW, Luangsa-ard JJ, Wingfield MJ et al (2018) Fungal planet description sheets: 785–867. Persoonia 41:238–417. https://doi.org/10.3767/persoonia.2018.41.12
Francis JO, Kirschenheuter GP, Ware GM et al (1988) Beta-cyclodextrin post-column fluorescence enhancement of aflatoxins for reverse-phase liquid chromatographic determination in corn. J Assoc Off Anal Chem 71:725–728
Gams W, Hawksworth DL (1975) Identity of Acrocylindrium oryzae Sawada and a similar fungus causing sheath-rot of rice. Kavaka 3:57–61
Gerlach W, Nirenberg H (1982) The genus Fusarium: a pictorial atlas. Biologische Bundesanstalt für Land- und Forstwirtschaft
Giraldo A, Gené J, Cano J et al (2012) Two new species of Acremonium from Spanish soils. Mycologia 104:1456–1465. https://doi.org/10.3852/11-402
Giraldo A, Gené J, Sutton DA et al (2015) Phylogeny of Sarocladium (Hypocreales). Persoonia 34:10–24. https://doi.org/10.3767/003158515X685364
Hara S, Fennell DI, Hesseltine CW (1974) Aflatoxin-producing strains of Aspergillus flavus detected by fluorescence of agar medium under ultraviolet light. Appl Microbiol 27:1118–1123
Jacobs K, Bergdahl DR, Wingfield MJ et al (2004) Leptographium wingfieldii introduced into North America and found associated with exotic Tomicus piniperda and native bark beetles. Mycol Res 108:411–418. https://doi.org/10.1017/S0953756204009748
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lefort V, Longueville J-E, Gascuel O (2017) SMS: smart model selection in PhyML. Mol Biol Evol 34:2422–2424. https://doi.org/10.1093/molbev/msx149
Liu X, Guo Z, Huang G (2017) Sarocladium brachiariae sp. nov., an endophytic fungus isolated from Brachiaria brizantha. Mycosphere 8:827–834. https://doi.org/10.5943/mycosphere/8/7/2
Mashima J, Kodama Y, Fujisawa T et al (2016) DNA Data Bank of Japan. Nucleic Acids Res 45:D25–D31. https://doi.org/10.1093/nar/gkw1001
O’Donnell K, Kistler HC, Cigelnik E et al (1998) Multiple evolutionary origins of the fungus causing Panama disease of banana: concordant evidence from nuclear and mitochondrial gene genealogies. Proc Natl Acad Sci 95:2044–2049. https://doi.org/10.1073/pnas.95.5.2044
Palleroni NJ (1984) Genus I. Pseudomonas Migula 1894. In: Bergey’s Manual of Systematic Bacteriology. Williams & Wilkins, Baltimore, pp 141–199
Perdomo H, Sutton DA, Garcia D et al (2011) Spectrum of clinically relevant Acremonium species in the United States. J Clin Microbiol 49:243–256. https://doi.org/10.1128/JCM.00793-10
Rivera-Varas VV, Freeman TA, Gudmestad NC et al (2007) Mycoparasitism of Helminthosporium solani by Acremonium strictum. Phytopathology 97:1331–1337. https://doi.org/10.1094/PHYTO-97-10-1331
Ronquist F, Teslenko M, van der Mark P et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Seifert KA (1985) A monograph of Stibella and some allied Hyphomycetes. Centraalbureau voor Schimmelcultures, Baarn
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stirling D (2003) Technical notes for sequencing difficult templates. In: Bartlett JMS, Stirling D (eds) PCR Protocols. Humana Press, Totowa, pp 401–402
Summerbell RC, Gueidan C, Schroers H-J et al (2011) Acremonium phylogenetic overview and revision of Gliomastix, Sarocladium, and Trichothecium. SIM 68:139–162. https://doi.org/10.3114/sim.2011.68.06
Summerbell RC, Gueidan C, Guarro J et al (2018) The protean Acremonium. A. sclerotigenum/egyptiacum: revision, food contaminant, and human disease. Microorganisms 6:88. https://doi.org/10.3390/microorganisms6030088
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577. https://doi.org/10.1080/10635150701472164
Voigt K, Wöstemeyer J (2000) Reliable amplification of actin genes facilitates deep-level phylogeny. Microbiol Res 155:179–195. https://doi.org/10.1016/S0944-5013(00)80031-2
White TJ, Bruns T, Lee S et al (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: PCR Protocols. Elsevier, Amsterdam, pp 315–322
Yeh Y-H, Kirschner R (2014) Sarocladium spinificis, a new endophytic species from the coastal grass Spinifex littoreus in Taiwan. Bot Stud 55:25. https://doi.org/10.1186/1999-3110-55-25
Funding
This study was supported by Council of Agriculture, Executive Yuan, grant nos. 107AS-8.4.1-BQ-B5 and 108AS-8.4.2-BQ-B1.
Author information
Authors and Affiliations
Corresponding author
Additional information
Section Editor: Hans-Josef Schroers
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 46.6 kb)
Rights and permissions
About this article
Cite this article
Ou, JH., Lin, GC. & Chen, CY. Sarocladium species associated with rice in Taiwan. Mycol Progress 19, 67–80 (2020). https://doi.org/10.1007/s11557-019-01543-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-019-01543-w